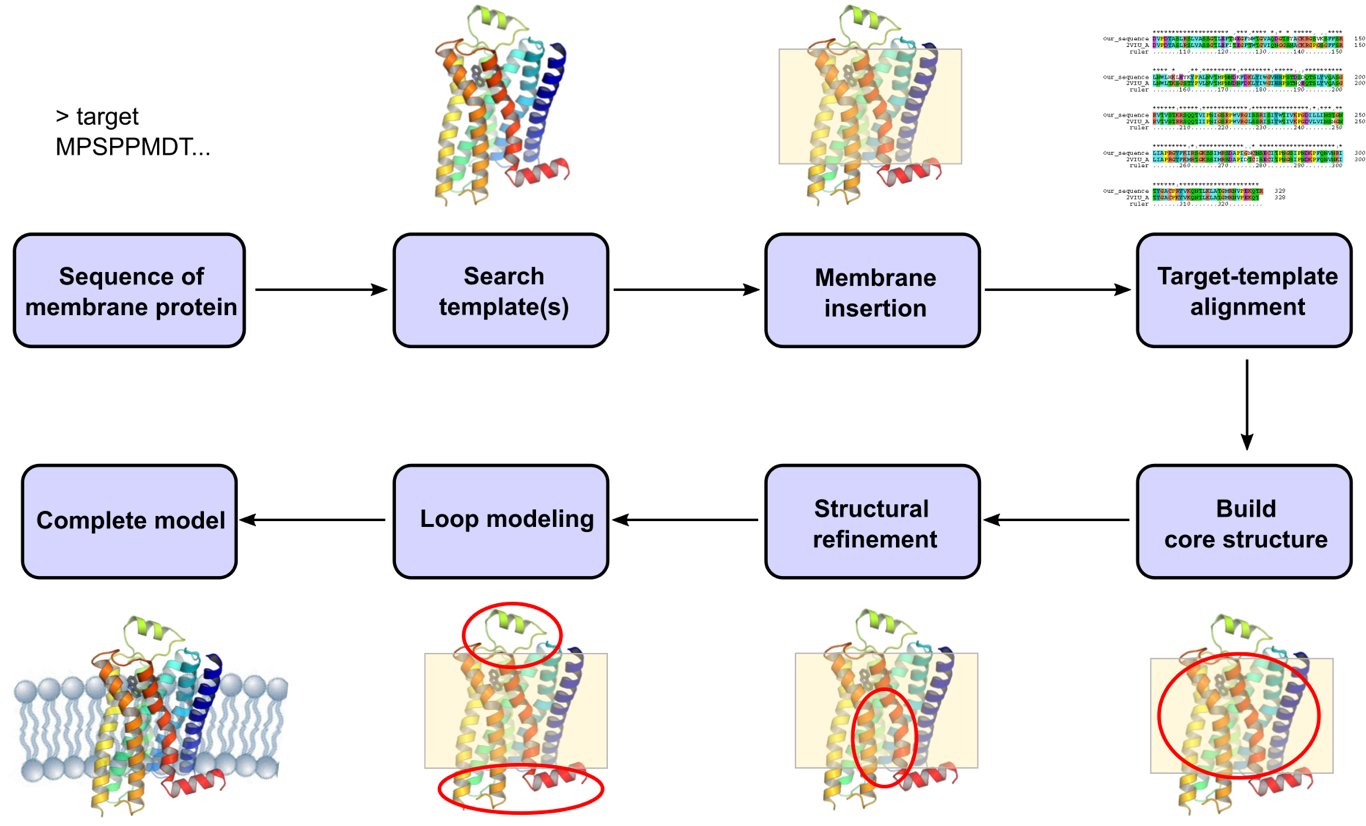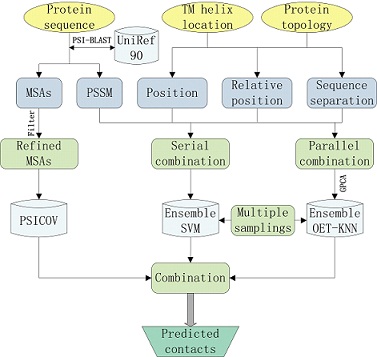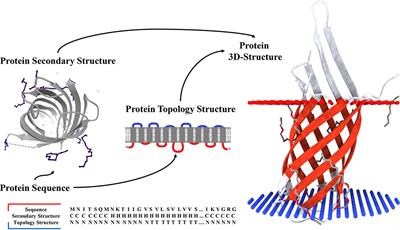
Frontiers | TMPSS: A Deep Learning-Based Predictor for Secondary Structure and Topology Structure Prediction of Alpha-Helical Transmembrane Proteins

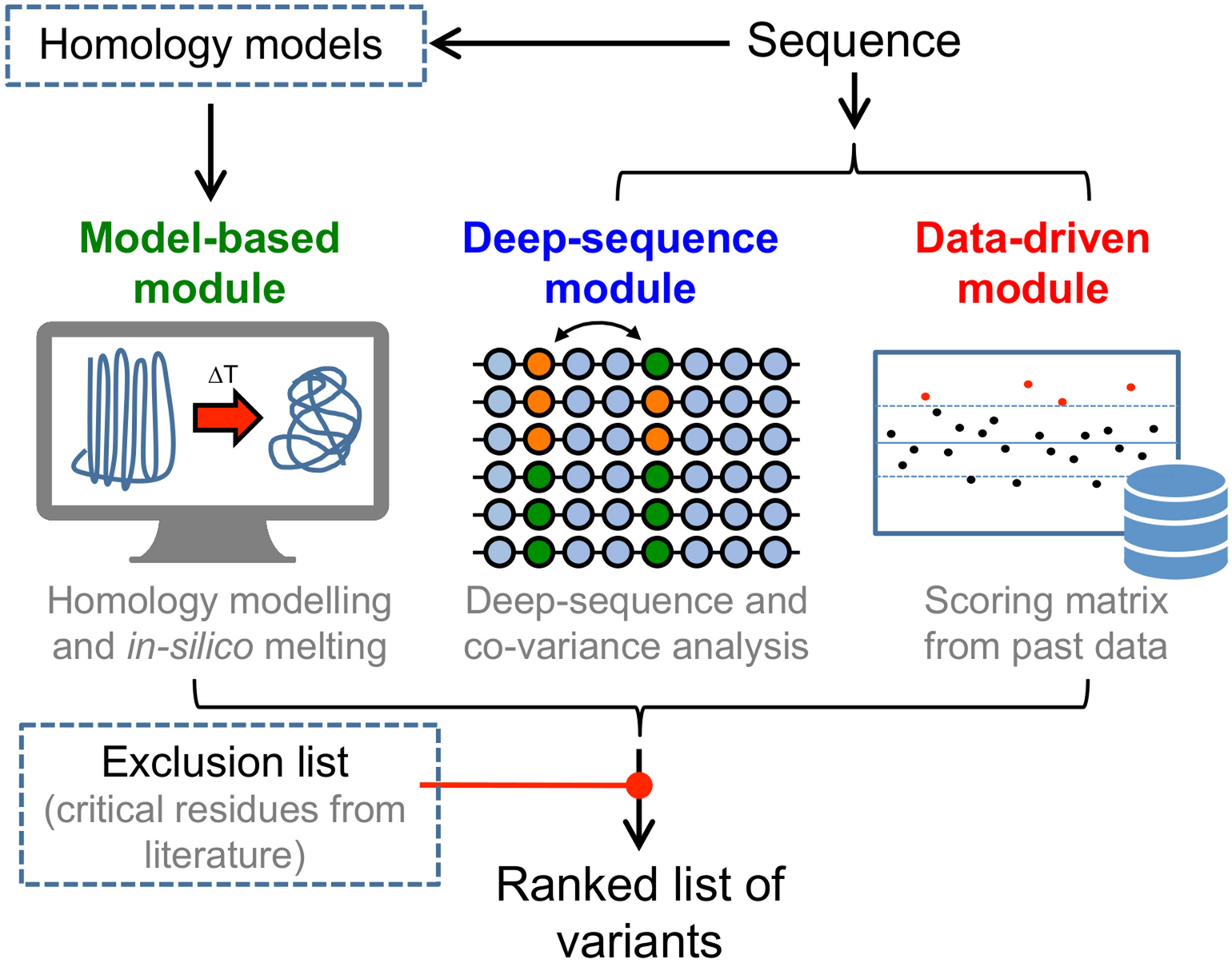
Machine learning in computational modelling of membrane protein sequences and structures: From methodologies to applications - ScienceDirect

Evolutionary analysis is used for predicting both the structure and... | Download Scientific Diagram

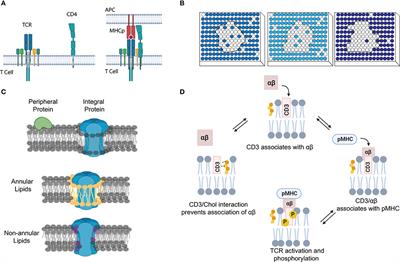
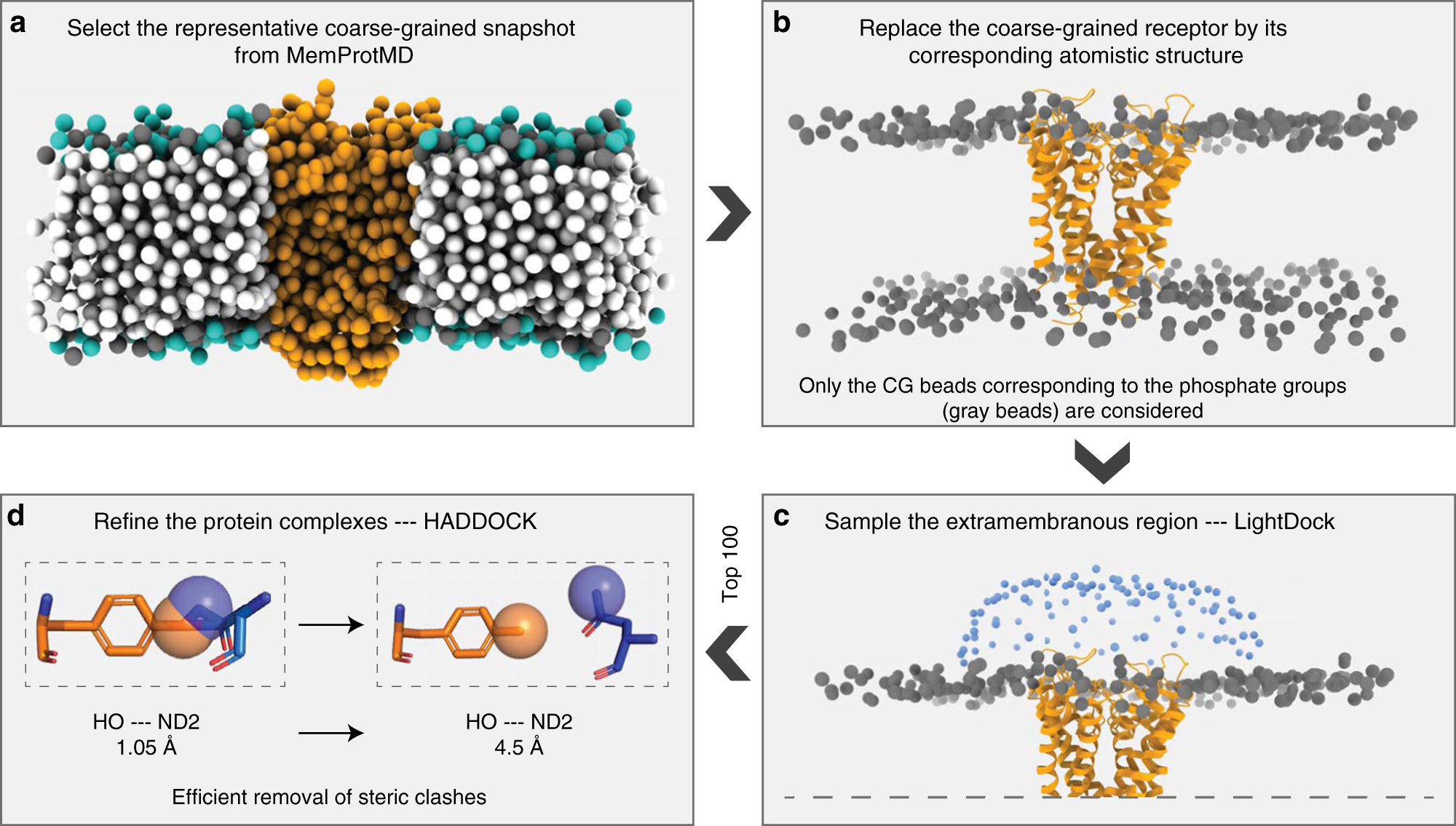
Getting to know each other: PPIMem, a novel approach for predicting transmembrane protein-protein complexes - ScienceDirect

The de novo design of a biocompatible and functional integral membrane protein using minimal sequence complexity | Scientific Reports
Membrane contact probability: An essential and predictive character for the structural and functional studies of membrane proteins | PLOS Computational Biology

Computational studies of membrane proteins: Models and predictions for biological understanding - ScienceDirect