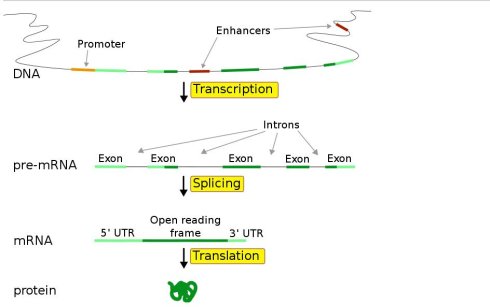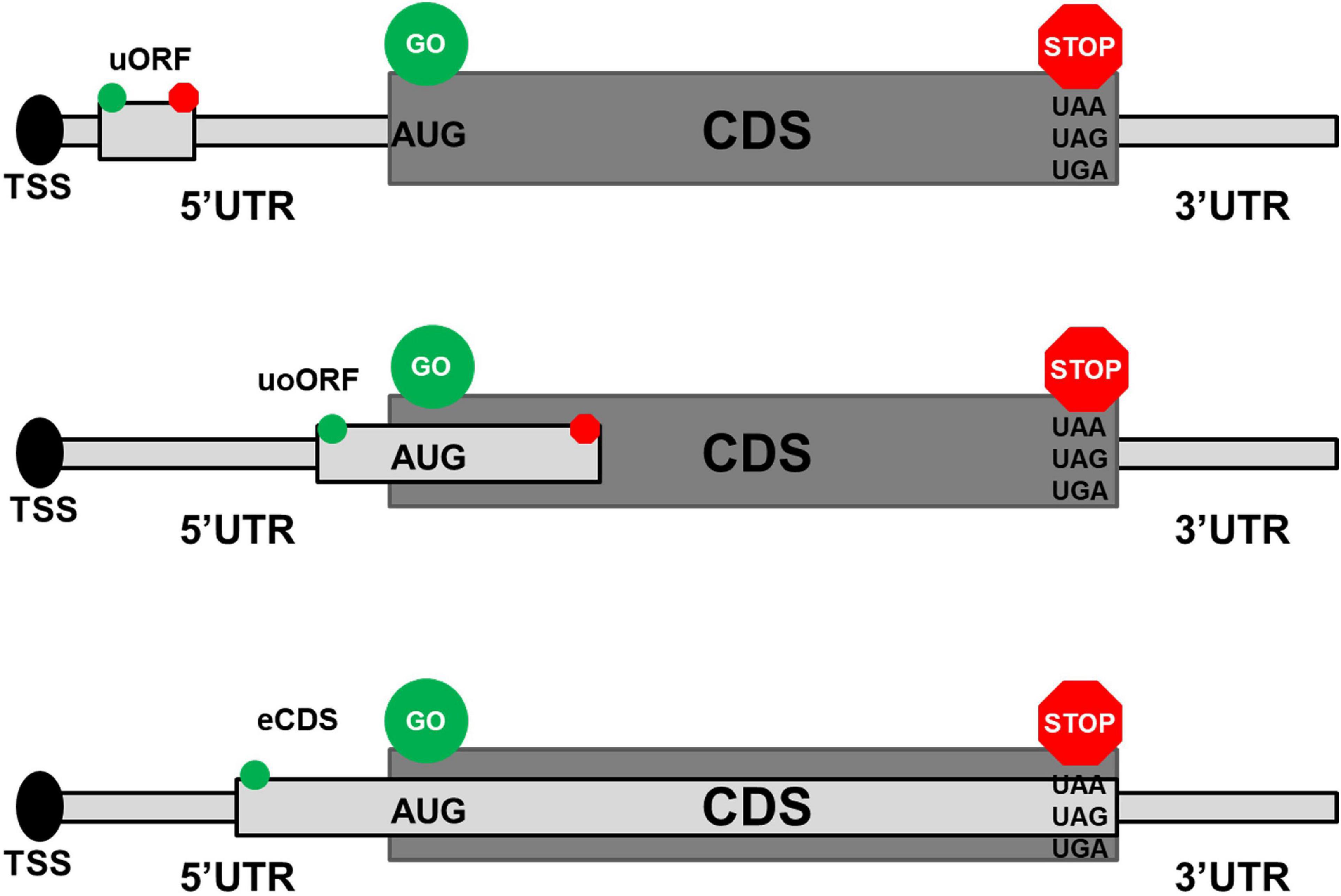
Frontiers | Common and Rare 5′UTR Variants Altering Upstream Open Reading Frames in Cardiovascular Genomics

DR4 and DR5 transcripts compared. CDS coding sequence, UTR untranslated... | Download Scientific Diagram

Widespread Differential Expression of Coding Region and 3′ UTR Sequences in Neurons and Other Tissues - ScienceDirect

Gene structure (5 0 UTR-CDS-3 0 UTR, including the corresponding number... | Download Scientific Diagram
Distinct expression of select and transcriptome-wide isolated 3'UTRs suggests critical roles in development and transition states | PLOS ONE

Cells | Free Full-Text | Protein Binding to Cis-Motifs in mRNAs Coding Sequence Is Common and Regulates Transcript Stability and the Rate of Translation

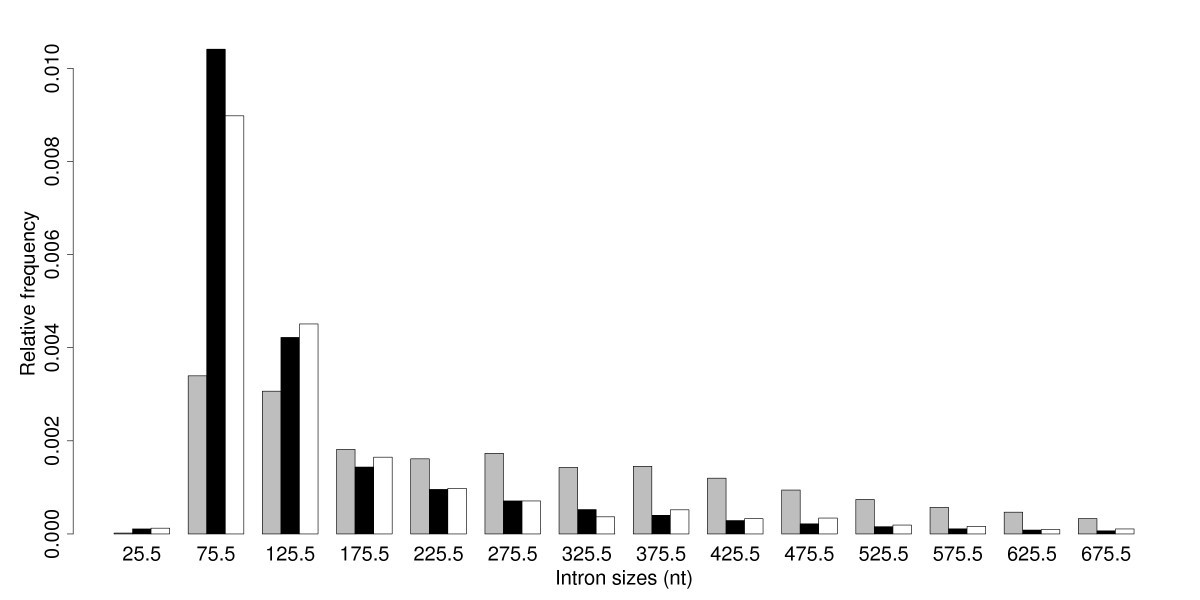
Length distributions of 5 UTR, 3 UTR and CDS introns (A) and position... | Download Scientific Diagram

Distinct expression of select and transcriptome-wide isolated 3'UTRs suggests critical roles in development and transition states | bioRxiv
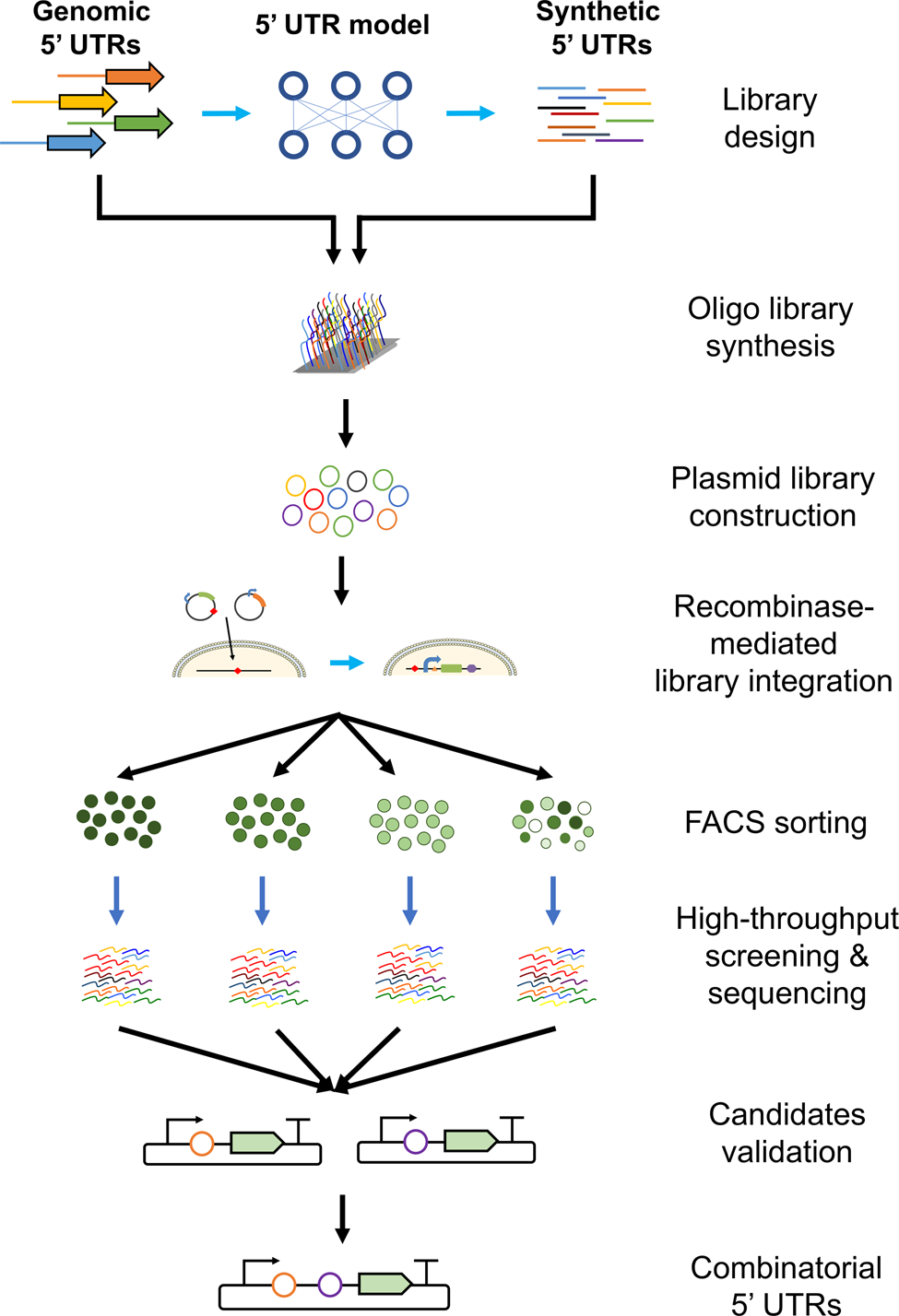
High-throughput 5′ UTR engineering for enhanced protein production in non-viral gene therapies | Nature Communications
GitHub - saketkc/gencode_regions: Extract 3'UTR, 5'UTR, CDS, Promoter, Genes, Introns, Exons from GTF files

Variation in upstream open reading frames contributes to allelic diversity in maize protein abundance | PNAS


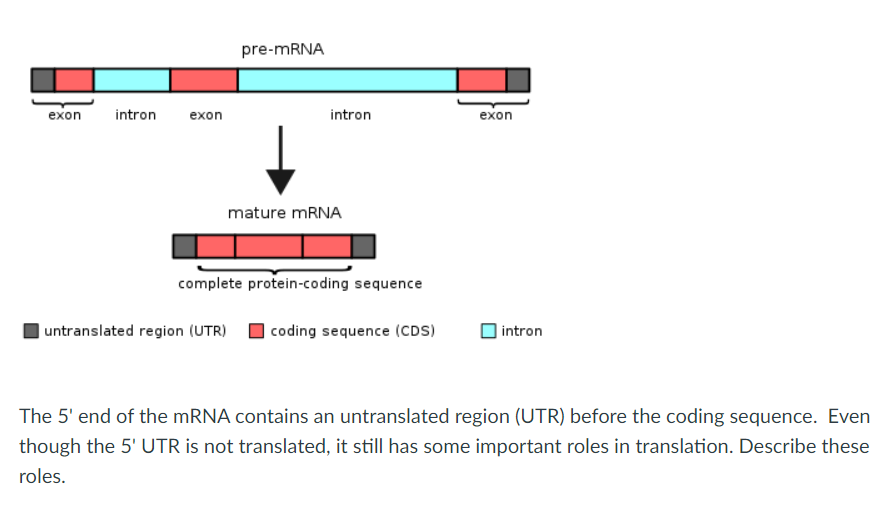
![미생물] ORF vs CDS, exon vs intron, splicing, UTR, 5'CAP, poly A tail 핵심 요약 정리 : 네이버 블로그 미생물] ORF vs CDS, exon vs intron, splicing, UTR, 5'CAP, poly A tail 핵심 요약 정리 : 네이버 블로그](https://mblogthumb-phinf.pstatic.net/MjAyMTA3MDJfMTc5/MDAxNjI1MjM0Mjc2NTU3.8knBWto8OCUDkdnQaY4KLjmo13I9yVK3TdySH9x9Zb0g.eZBwMhrtBboUXyk--cmvFfalanteQZ_3-Fs8gkqExQcg.PNG.wkdalgus94/image.png?type=w800)